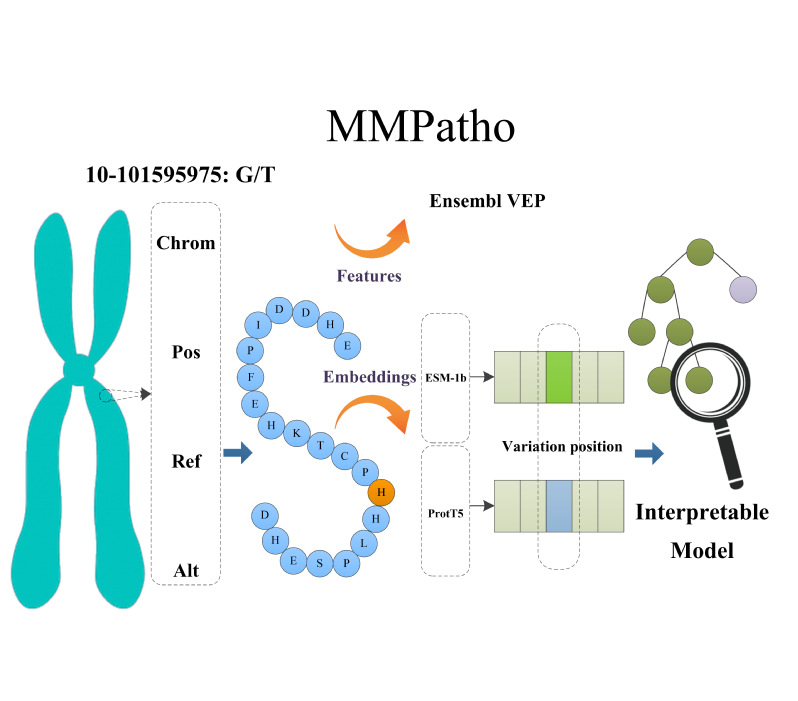
 | MMPatho
Category: [Genome-wide variants]
Description: Interpretable model for missense mutation pathogenic prediction and annotation.
Journal: Submitted to JCIM.
| 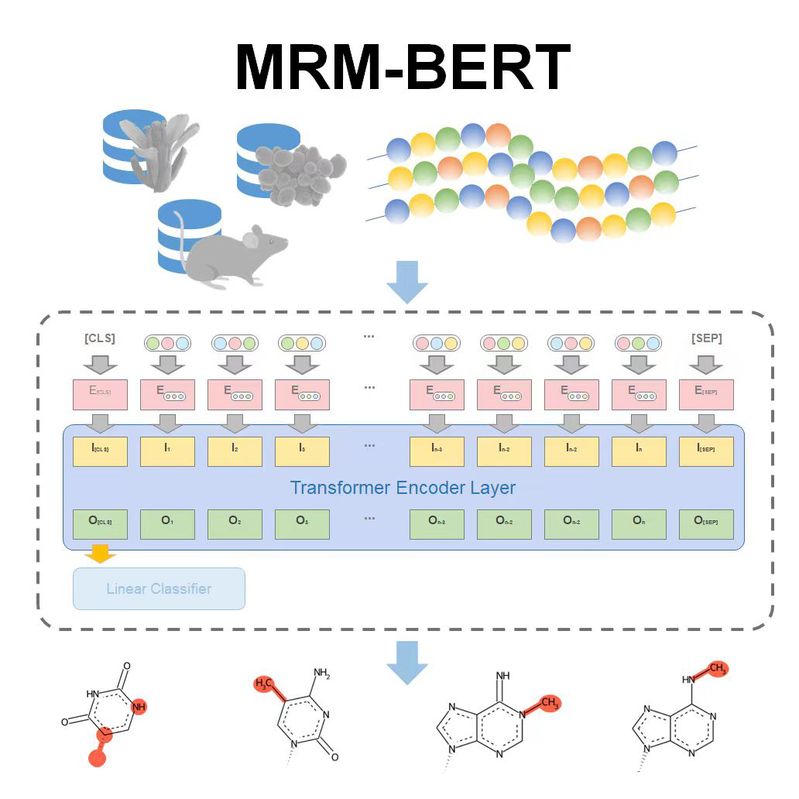 | MRM-BERT (Backup)
Category: [RNA Modification Prediction]
Description: Predicting RNA modification using biological language model.
Journal:IEEE TCBB. |
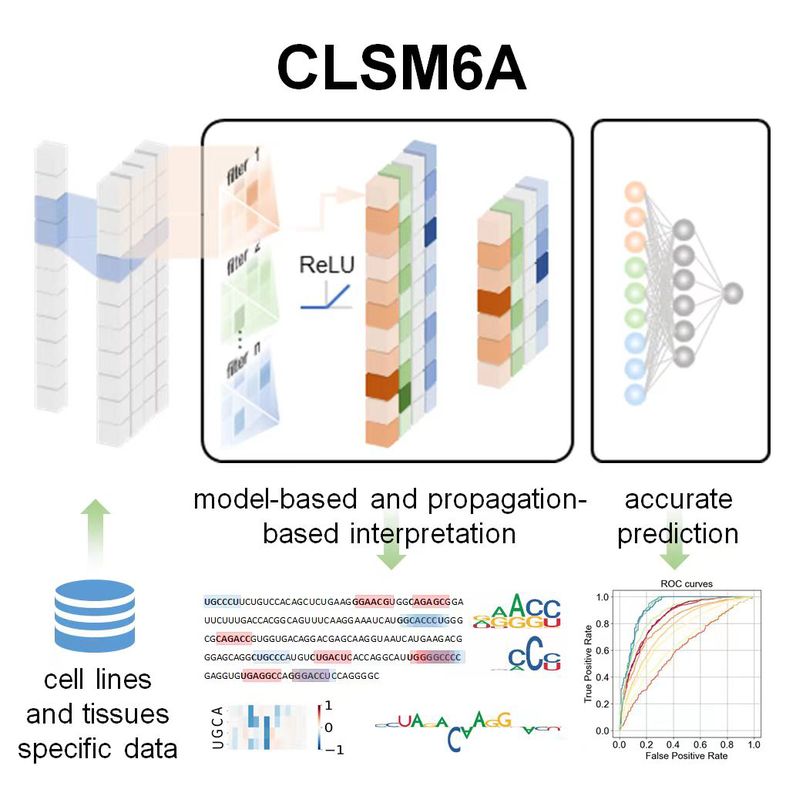 | CLSM6A
Category: [m6A RNA]
Description: Interpreting prediction models to reveal genetic insights of m6A.
Journal: Submitted to Briefings in Bioinformatics.
|  | MRM-BERT
Category: [RNA Modification Prediction]
Description: Predicting RNA modification using biological language model.
Journal:IEEE TCBB. |
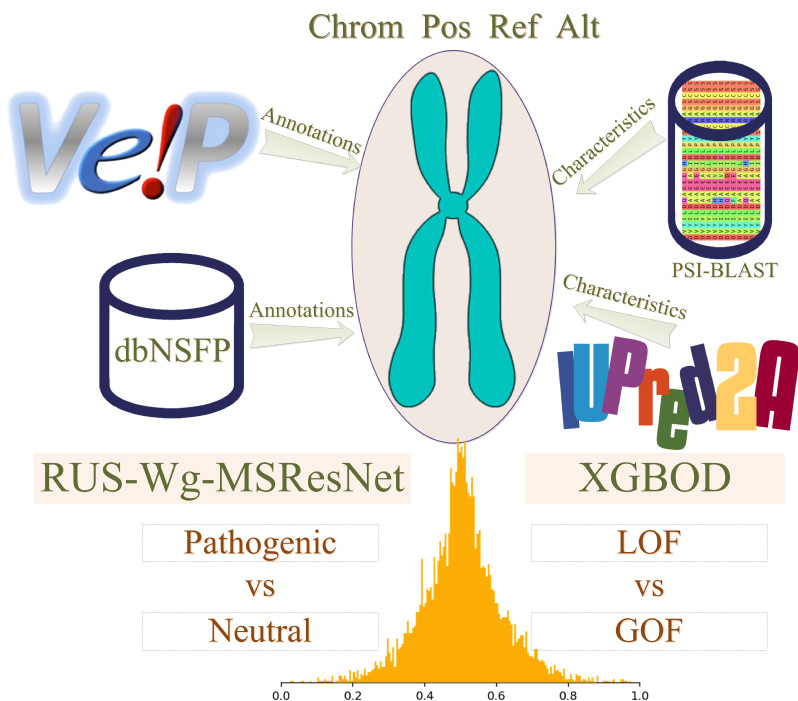 | VPatho
Category: [Genome-wide variants]
Description: Prediction of pathogenicity and function impact of variants.
Journal: Briefings in Bioinformatics.
| 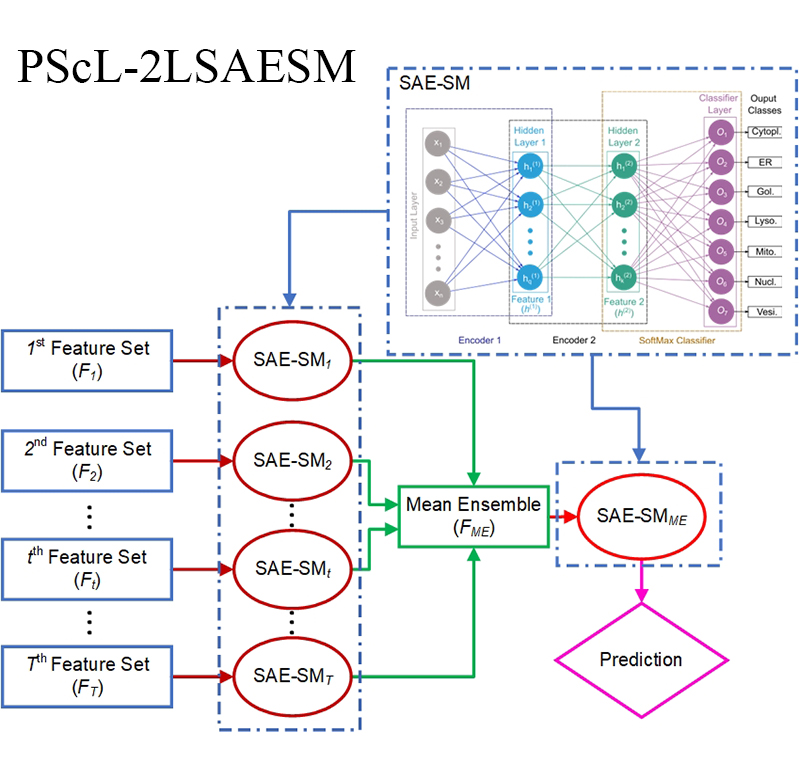 | PScL-2LSAESM
Category: [Protein subcellular localization]
Description: Predicting protein subcellular localization using two-level and mean sensemble method.
Journal:Bioinformatics. |
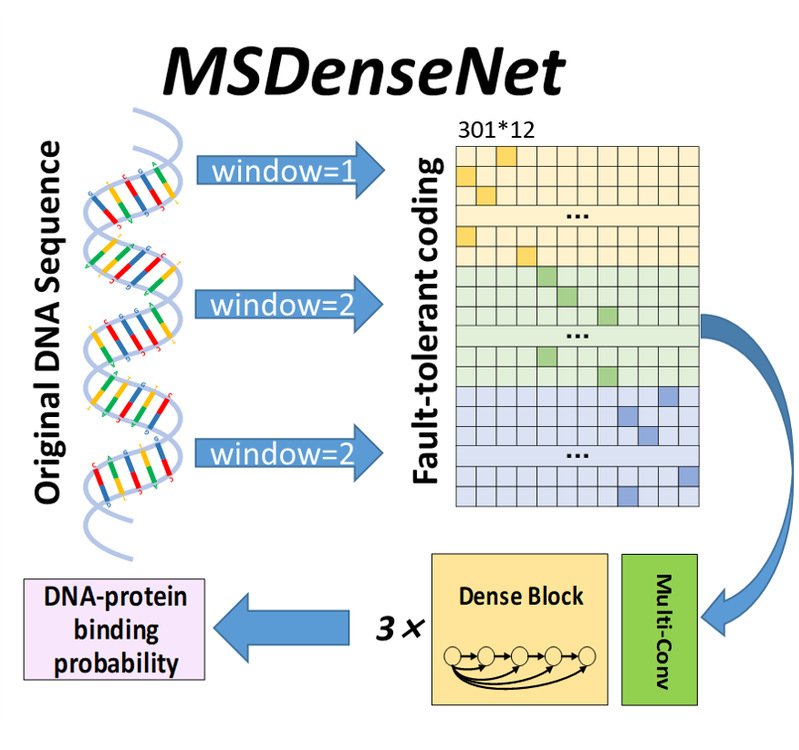 | MSDenseNet
Category: [DNA-protein]
Description: Prediction of DNA-protein interactions by using multi-scale dense CNN.
Journal: Analytical Biochemistry.
| 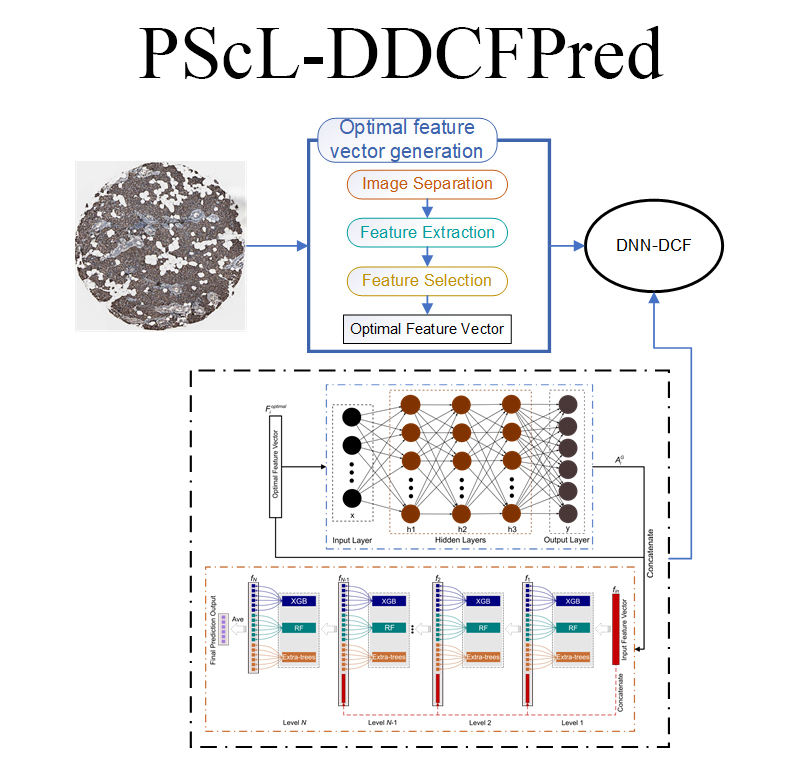 | PScL-DDCFPred
Category: [Protein subcellular localization]
Description: Predicting protein subcellular localization using deep learning technology.
Journal:Bioinformatics. |
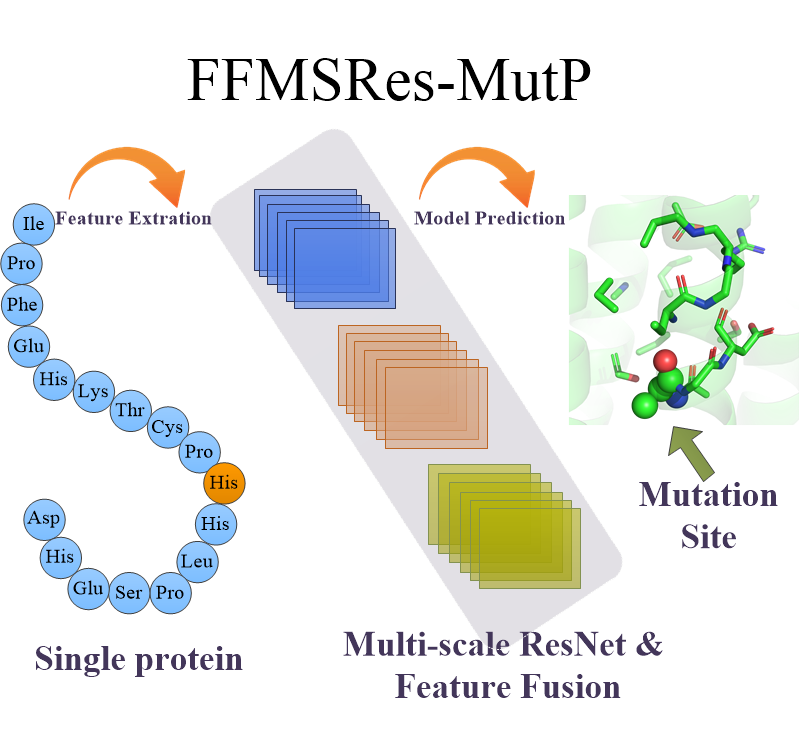 | FFMSRes-MutP
Category: [Mutations prediction]
Description: Prediction of disease-related nsSNPs by integrating deep feature fusion and multi-scale ResNet.
Journal: Briefings in Bioinformatics.
| 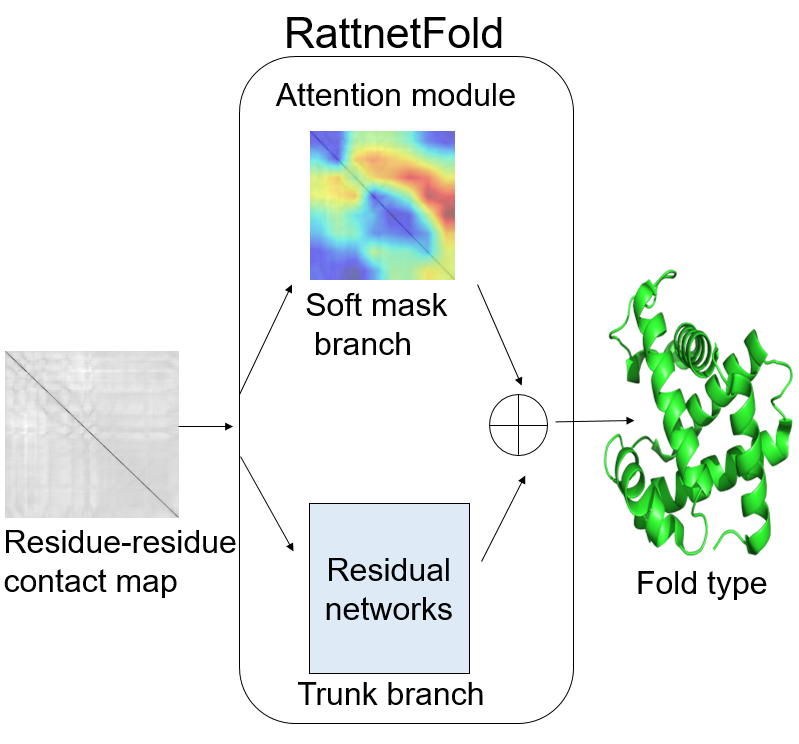 | RattentFold
Category: [Protein foldprediction]
Description: Predicting protein foldusing residual network
Journal:Analytical Biochemistry. |
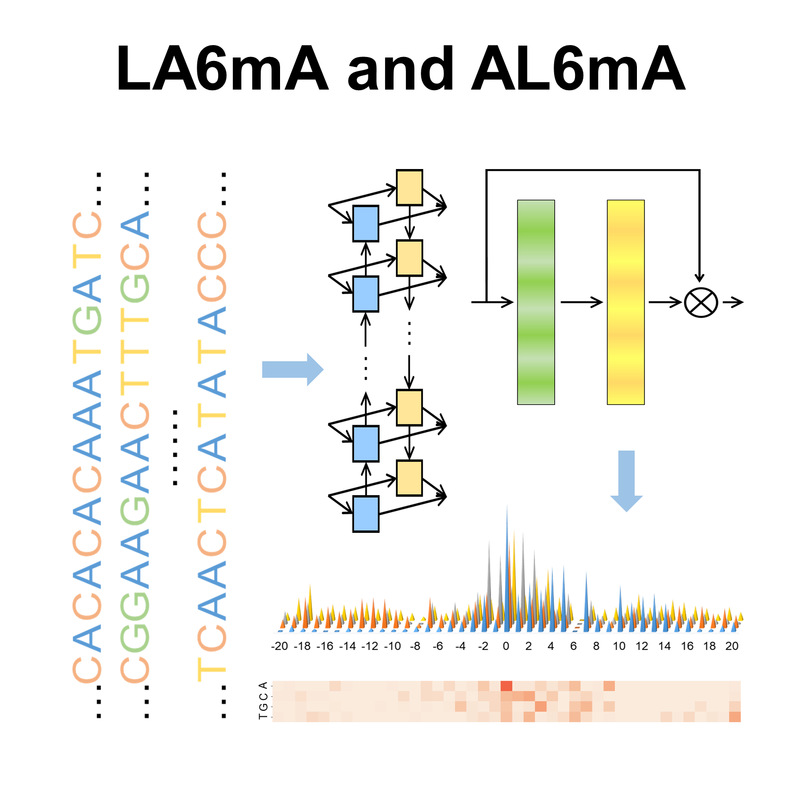 | AL6mA
Category: [DNA N6-methyladenine]
Description: Leveraging the attention mechanism to improve the identification of DNA N6-methyladenine sites.
Journal: Briefings in Bioinformatics.
| 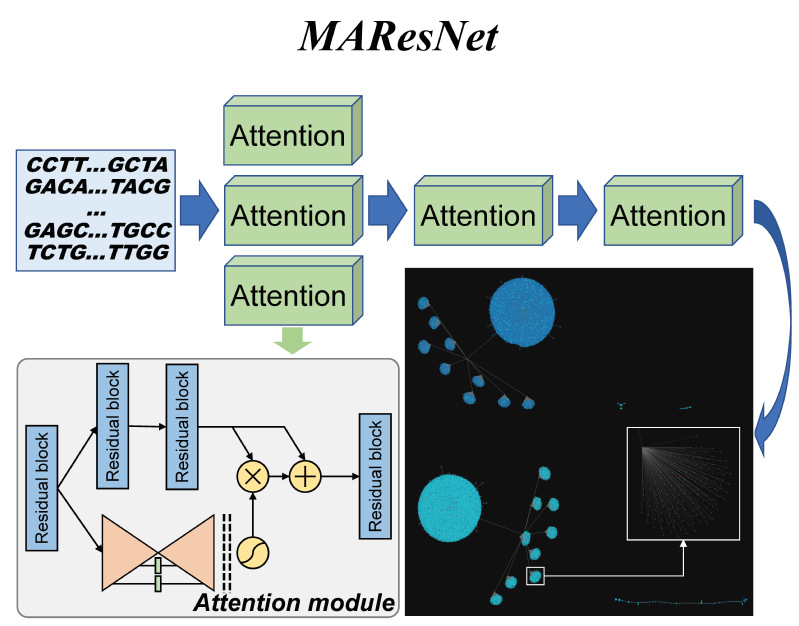 | MAResNet
Category: [TF binding sites prediction]
Description: Predicting transcription factor binding sites by combining attention mechanism and residual network
Journal:Briefings in Bioinformatics. |
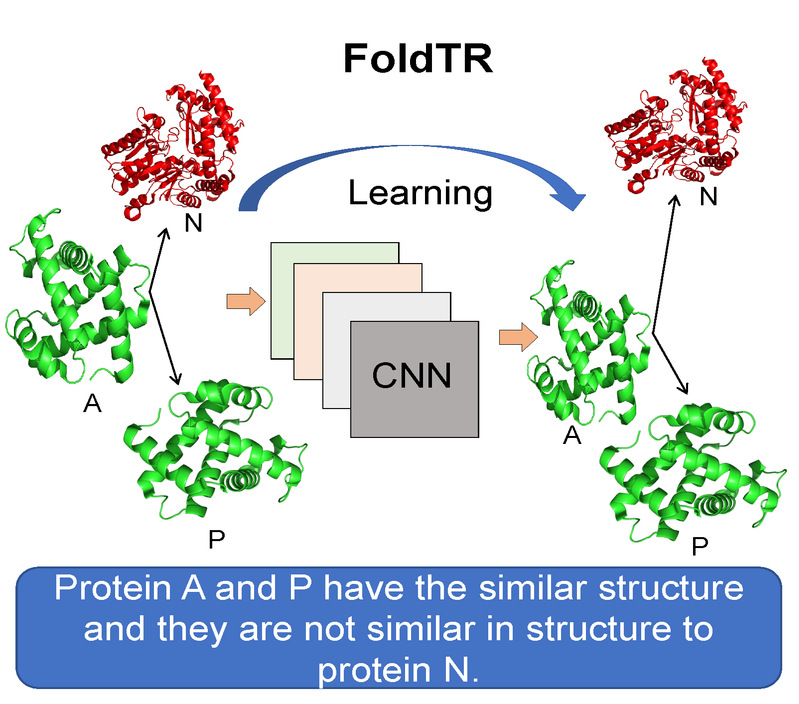 | FoldTR
Category: [Protein fold recognition]
Description: Protein fold recognition recognition using ensemble deep learning.
Journal: Briefings in Bioinformatics. | 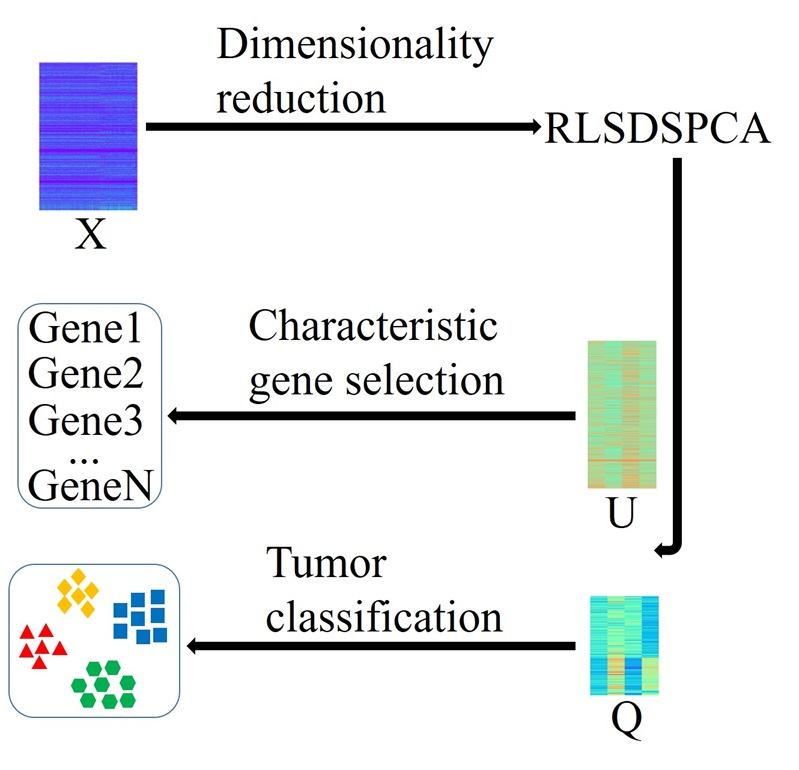 | RLSDSPCA
Category: [Characteristic Gene Selection]
Description: Characteristic Gene Selection and Tumor Classification by RLSDSPCA
Journal:Briefings in Bioinformatics. |
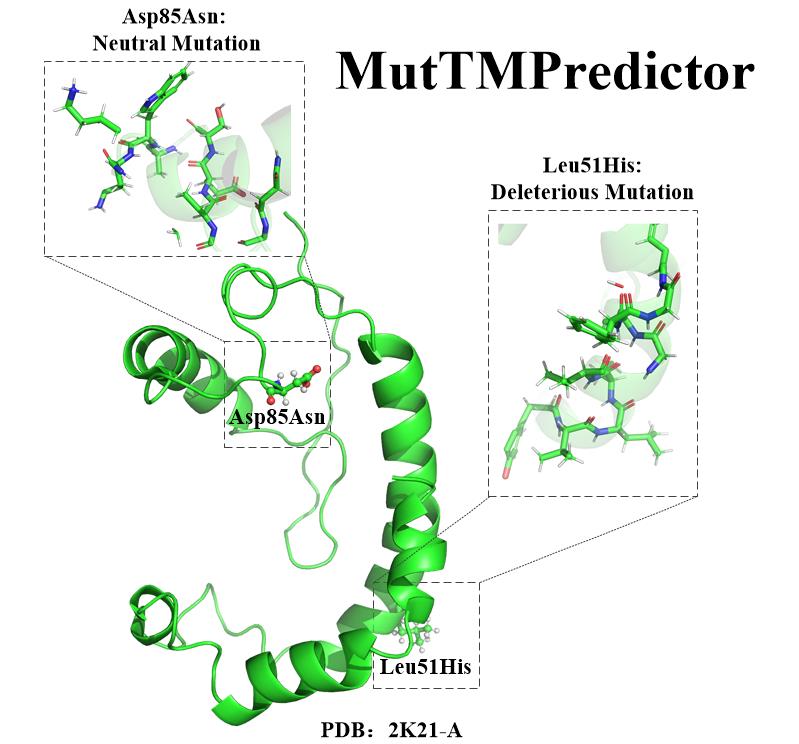 | MutTMpredictor
Category: [Mutations prediction]
Description: Disease-related mutations prediction in transmembrane protein.
Journal: Computational and Structural Biotechnology Journal. | 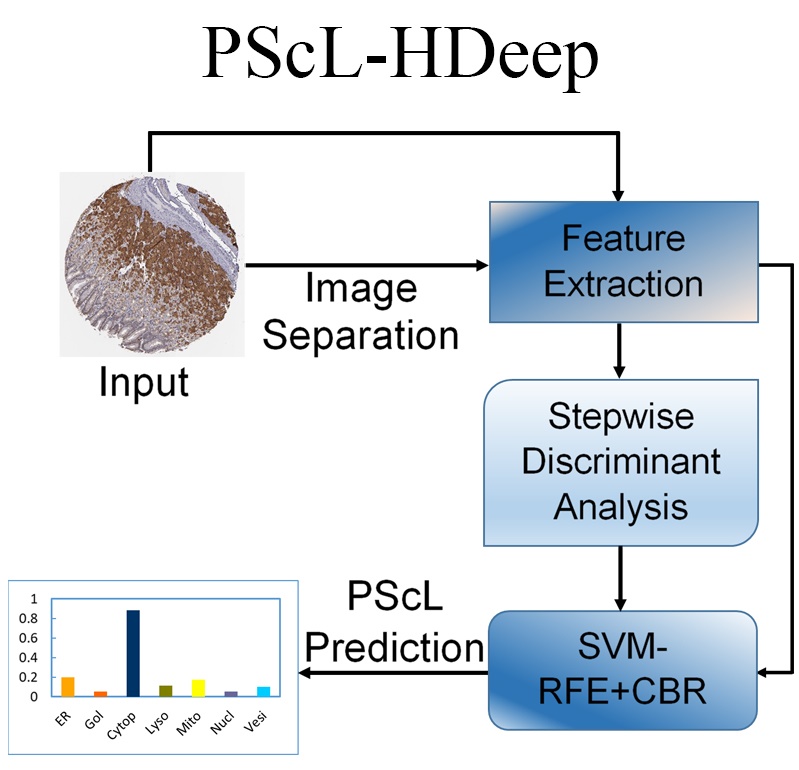 | PScL-HDeep
Category: [Protein subcellular localization]
Description: Image-based prediction of protein subcellular location in human tissue.
Journal:Briefings in Bioinformatics. |
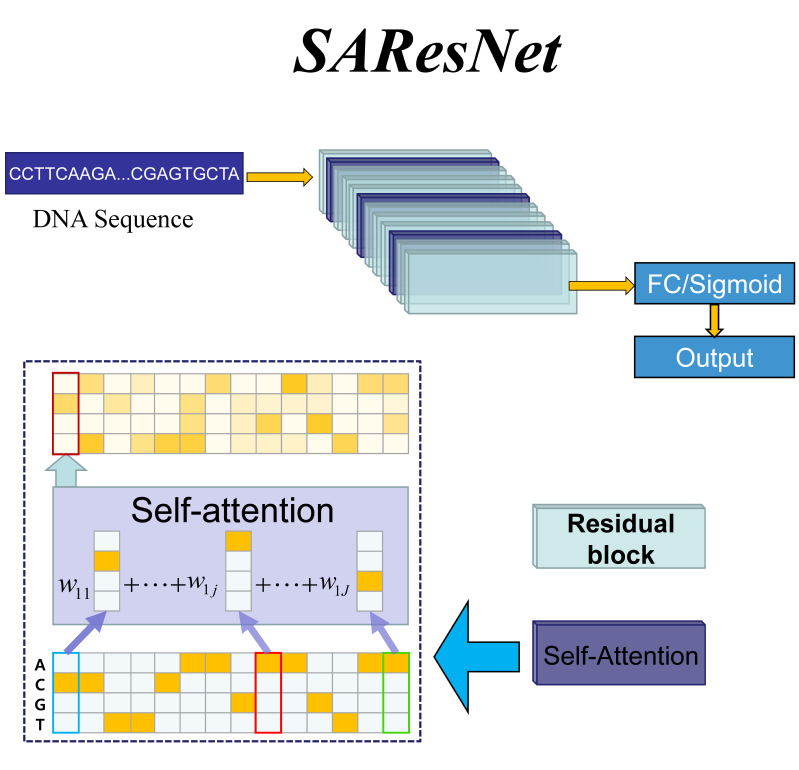 | SAResNet
Category: [DNA-Protein binding]
Description: Predicting DNA-Protein binding.
Journal: Briefings in Bioinformatics |  | VGGfold
Category: [Protein Fold Recognition]
Description: Visualizing protein fold recognition.
Journal:Briefings in Bioinformatics |
 | DCFCrystal
Category: [Protein Crystallization]
Description: Predicting Protein Crystallization.
Journal: Briefings in Bioinformatics | 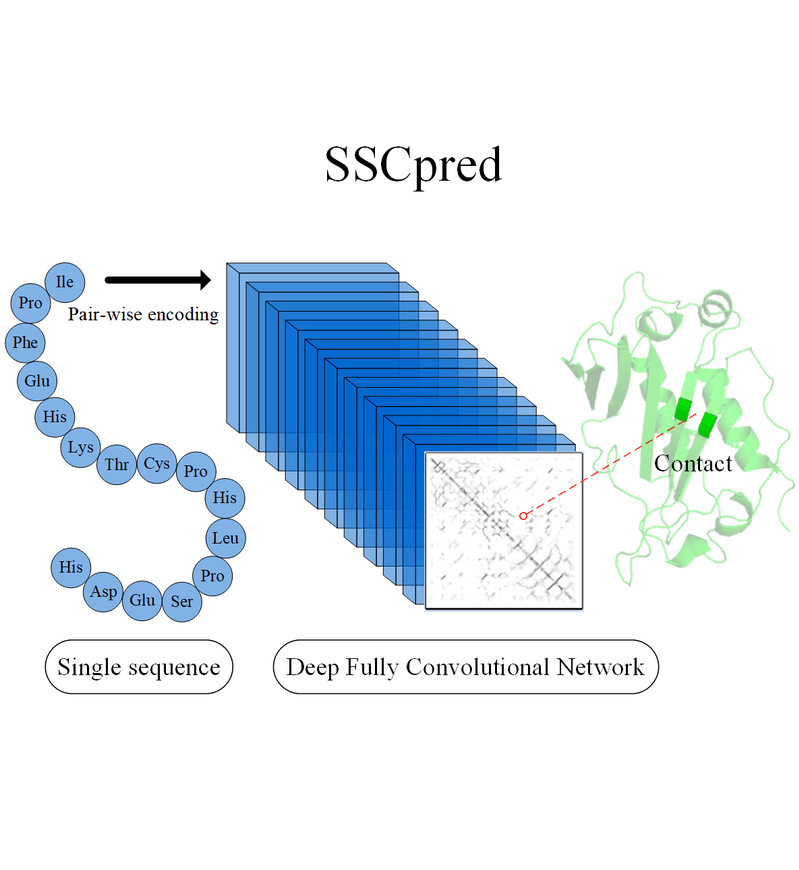 | SSCPred
Category: [Protein Contact Map]
Description: TargetingATP&ADP binding sitesfrom protein Sequences.
Journal: JCIM |
 | DNAPred
Category: [DNA-Binding Sites]
Description: Predicting DNA-Binding
Journal: JCIM |  | ATP-ADP
Category: [Protein-Ligand Interactions]
Description: TargetingATP&ADP binding sitesfrom protein Sequences.
Journal: ACPR |
 | TargetDBP
Category: [DNA-Binding Protein]
Description: Predicting DNA-Binding proteins.
Journal: IEEE TCBB |  | TargetDBP+
Category: [DNA-Binding Protein]
Description: Predicting DNA-Binding proteins.
Journal: IEEE TCBB |
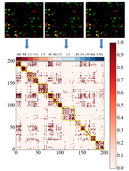 | SSC-LRR
Category: [Gene Expression Data]
Description: Cancer Classification on Gene Expression Data.
Journal: IEEE TCBB | 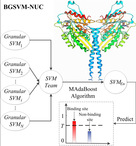 | BGSVM-NUC
Category: [Protein-Nucleotide interactions]
Description: Targeting protein-nucleotide interactions from protein
Journal: CCHT |
 | TargetM1A
Category: [N1-methyladenosine sites]
Description: Identifying m1A Sites from RNA Sequences.
Journal:
Paper download(unavailable) |  | TargetDNA
Category: [DNA-protein interactions]
Description: Targeting DNA-protein interactions from protein Sequences.
Journal: IEEE TCBB |
| TargetM6A
Category: [N6-methyladenosine sites]
Description: Identifying m6A Sites from RNA Sequences via Position-specific Nucleotide Propensity.
Journal:IEEE TNB | 
| M6A-HPCS
Category: [N6-methyladenosine sites]
Description: Targeting m6A sites from RNA sequences based on the selection of nucleotide physical-chemical properties.
Journal: Analytical Biochemistry |
| SSWRF-PPI
Category: [Protein-Protein Interaction Sites]
Description: Protein-Protein interaction sites prediction by ensembling SVM and sample-weighted random forests.
Journal: Neurocomputing | 
| TargetNUCs
Category: [Protein-Nucleotide Interaction]
Description: Dynamic Query-Driven Sample Rescaling Strategy for targeting protein-nucleotide binding sites.
Journal: Neurocomputing |
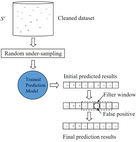
| DC-RF-RUS-PF
Category: [Protein-Protein Interaction Sites]
Description: Prediction of Protein-Protein Interaction Sites with Machine Learning based Data-Cleaning and Post-Filtering Procedures.
Journal: JMBI | 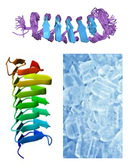
| TargetFreeze
Category:[Antifreeze Protein Prediction]
Description: Antifreeze Protein Prediction by Exploring Multiple Sequential Features and Support Vector Machine.
Journal: JMBI |
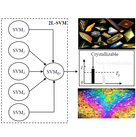
| TargetCrys Category: [Protein Crystallization Prediction]
Description: Protein Crystallization Prediction by Fusing Multi-View Features with Two-Layered SVM.
Journal: Amino Acids |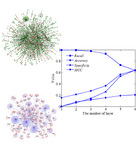 
| CRF-PPI
Category:[Protein-Protein Interactions]
Description: A Cascade Random Forests Algorithm for Predicting Protein-Protein Interaction Sites.
Journal: IEEE TNB |
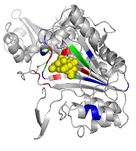
| OSML
Category: [Protein-Nucleotide Interaction]
Description: A Query-Driven Dynamic Machine Learning Model for Predicting Protein-Ligand Binding Sites.
Journal: IEEE TNB | 
| TargetSOS
Category: [Protein-Nucleotide Interaction]
Description: A supervised over-sampling (SOS) predictor for targeting protein-nucleotide binding residues
Journal: PLoS ONE |

| TargetGDrug
Category: [Protein-Drug Interaction]
Description: Predicting GPCR-Drug Interactions by Random Forest with Drug Association Matrix Based Post-processing Procedure.
Journal: CBAC | 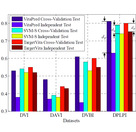
| TargetVita
Category: [Protein-Vitamin Interaction]
Description: Enhancing Protein-Vitamin Binding Residues Prediction by Multiple Heterogeneous Subspace SVMs Ensemble.
Journal: BMC Bioinformatics |
| TargetDisulfide
Category:[Disulfide Connectivity Prediction]
Description: Disulfide Connectivity Prediction Based on Modelled Protein 3D Structural Information and Random Forest Regression.
Journal: IEEE TCBB | 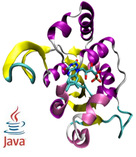
| TargetATP
Category: [Protein-ATP Interaction]
Description: A Java Web Start Application for Protein-ATP Binding Residue Prediction.
Journal: Neurocomputing |
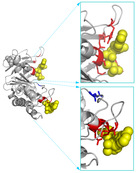
| TargetATPsite
Category:[Protein-ATP Interaction]
Description: A Template-free Method for ATP Binding Sites Prediction with Residue Evolution Image Sparse Representation.
Journal: JCC | 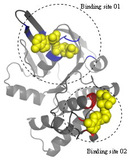
| TargetS
Category: [Protein-Nucleotide Interaction]
Description: A ligand-specific template-free predictor for targeting protein-ligand binding sites.
Journal: IEEE TCBB |
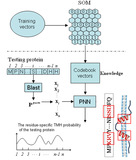 | SONPNN
Category: [Helices Prediction]
Description: An Efficient Non-Parametric Model for Predicting Transmembrane Helices.
Journal:Amino Acids |  | MPSP
Category: [Subcellular Localization Prediction]
Description: Membrane Protein Subcellular Localization Prediction by Parallel Fusion of Multi-View Features.
Journal:IEEE TNB |
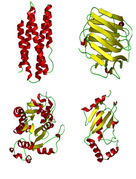
| COMSPA
Category: [Protein Structural Classes Prediction]
Description:Predicting Protein Structural Classes From Primary Sequence by Learning Multi-View Features in Complex Space.
Journal:Amino Acids | 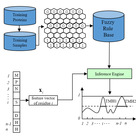
| SOMRuler
Category: [Helices Prediction]
Description: A Novel Interpretable Transmembrane Helices Predictor.
Journal: IEEE TNB |
| ULDNA
Category: [DNA-Binding Sites]
Description:Predicting DNA-Binding.
Journal:Submitted to Briefings in Bioinformatics |
|
|